Textbook Theory on Bacteria Challenged
Gene “Switch” Rule Not Universal, Study Finds
Discovery May Aid Smarter Antibiotics
Kolkata: For almost 50 years, biology textbooks around the world have taught a clear and confident rule about how bacteria switch their genes on. That rule is now being reconsidered — thanks in part to a team of Indian scientists.
Researchers from the Bose Institute, working with collaborators at Rutgers University, have found that a long-accepted model of bacterial gene control does not apply to all bacteria. Their study, published in the Proceedings of the National Academy of Sciences, challenges what has been considered a central textbook concept in microbiology.
To understand the discovery, think of a gene as a set of instructions inside a bacterial cell. To read those instructions, the cell uses a molecular machine called RNA polymerase. For decades, scientists believed this machine needs a helper molecule — known as a sigma factor — to begin reading a gene. The traditional view, based largely on research on Escherichia coli, suggested that the sigma factor attaches to RNA polymerase to start the process, then detaches once gene copying is underway. This repeated attaching and detaching became known as the “sigma cycle.”
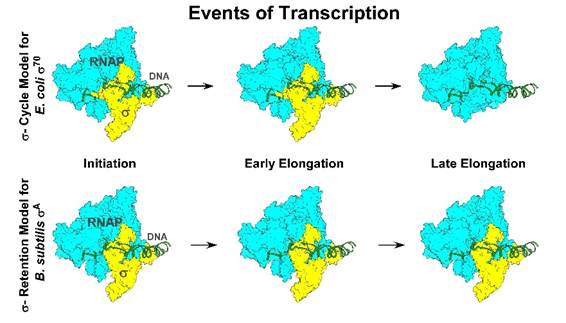
But the new research shows that this cycle is not universal. The team discovered that in another widely studied bacterium, Bacillus subtilis, the main sigma factor — called sigma A — does not leave after starting the gene-reading process. Instead, it remains firmly attached to RNA polymerase throughout the entire process of gene copying.
The scientists also examined a modified version of the E. coli sigma factor, one missing a small portion known as region 1.1. This altered version behaved like sigma A in Bacillus subtilis — it stayed attached. In contrast, the full-length E. coli sigma factor was found to detach randomly during gene copying, just as the traditional model suggests.
In simple terms, what was once believed to be a universal rule turns out to be only part of the story.
Using advanced laboratory techniques — including biochemical tests, chromatin immunoprecipitation, and fluorescence imaging that allowed them to observe these molecular events almost in real time — the researchers tracked how sigma factors behaved inside living systems. The direct evidence they gathered shows that the “sigma cycle” does not apply across all bacterial species.
Dr Jayanta Mukhopadhyay of the Bose Institute, a corresponding author of the study, said the findings fundamentally change how scientists think about bacterial gene regulation. Co-author Aniruddha Tewari added that the results open new avenues for understanding not only how bacteria control their genes today, but also how these control systems may have evolved over time.
The implications could extend far beyond academic debate. A clearer picture of how bacteria regulate their genes could help scientists design more effective antibiotics or regulatory inhibitors that block infection mechanisms. It may also help researchers engineer bacteria more efficiently to produce biofuels, biodegradable plastics, or therapeutic compounds.
The study was authored by Aniruddha Tewary, Shreya Sengupta, Soumya Mukherjee and Nilanjana Hazra from the Bose Institute, along with Yon W. Ebright, Richard H. Ebright and Jayanta Mukhopadhyay from Rutgers University.
In science, even long-standing rules are open to revision when new evidence appears. This study is a reminder that biology is still full of surprises — and that Indian laboratories are helping reshape our understanding of life at its most fundamental level.
– global bihari bureau





